Research Overview
Role of p53 in Cancers
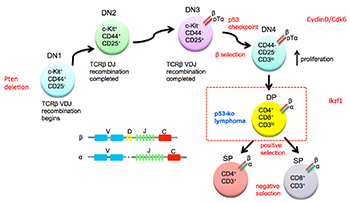
P53 is a tumor suppressor whose function is crucially important in guarding genome stability. It is the most commonly mutated gene found in fifty percent of all human cancers. We study the role of p53 in tumorigenesis in Li-Fraumeni Syndrome (LFS) and p53 knockout mouse models. Loss of p53 is associated with increased genome instability in cancers but whether p53 plays a similar role in the germline is not well understood. We studied families with LFS and showed no evidence for germline genome instability by genome sequencing. We created a genetic model termed, genetic regression, which explains the apparent observation of genetic anticipation and heterogeneity in LFS. We studied the evolution of thymic lymphoma in p53 knockout mice and identified the sequence of additional mutations required for tumorigenesis. We performed similar studies for liposarcoma in a mouse model. We continue to study LFS through sequencing of tumors from LFS individuals as well as tumors from LFS mouse models in different genetic hybrid backgrounds.
Cancer Genomics and Evolution
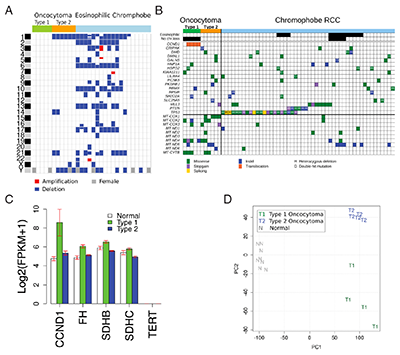
Cancers result from an evolutionary process in which mutations that inactivate tumor suppressors or activate oncogenes that lead to increased proliferation are selected for in cells undergoing tumorigenesis. We apply computational methods to identify tumor suppressors and oncogenes in various cancer types as well as provide a molecular classification of tumor types. Some of the tumor types we are studying include oncocytomas and neuroendocrine tumors where very little is known about the etiology of the disease especially the key mutations responsible for their tumorigenesis. We are characterizing molecular subtypes of these tumors by their transcriptional and mutation profiles. We are also identifying oncogenes and tumor suppressors in these cancers as well as the genetic pathways aberrantly perturbed. In acute myeloid leukemia and non-small cell lung cancer, we are studying the genetic, epigenetic and immunologic underpinnings of cancer evolution through treatment.
Gene Regulation
There are over a thousand different cell types in a human and the diverse phenotypes are determined by gene regulation. We use computational genomics and systems biology to study transcriptional and post-transcriptional gene regulation including large and small non-coding RNAs and repetitive elements.

